with 20 more rows Centre the dataįor PCA to work properly, you have to subtract the mean from each of the data dimensions. Thus, we might expect there to be little to no correlation between the variables they are uncorrelated. We draw two variables $x$ and $y$ from different random normal distributions. Let’s keep it simple and generate our own data we use just two dimensions so that we can visualise what’s happening.
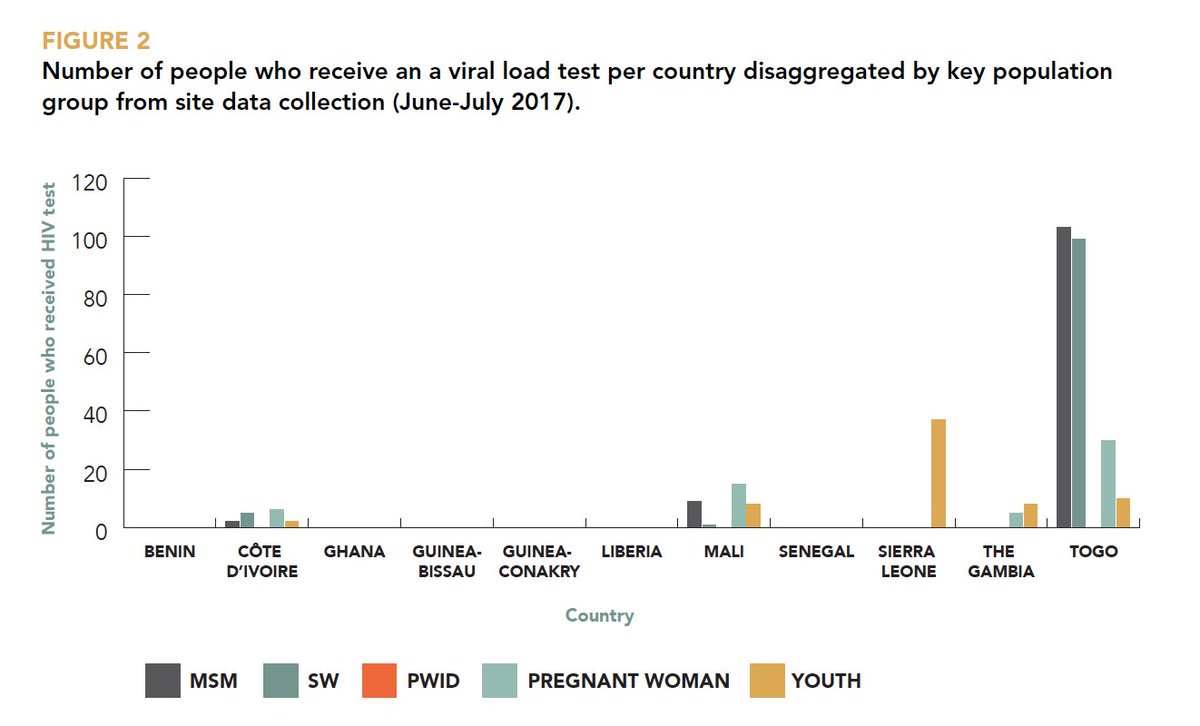
We demonstrate PCA by comparing the outcomes of the methodology on: Given that we are only interested in variance and not interested in the scale, the variable should be centered to have mean zero prior to PCA. PCA does this by finding features that show as much variation across observations or, put another way, PCA seeks a small number of dimensions that are as interesting as possible, where the concept of interesting is determined by how much the observations vary along each dimension. Imagine the explanatory variables or features of your data contain some redundant information (information that is captured by another feature) then there is the potential to summarise the same data with fewer new features. We paraphrase the excellent stack overflow answer: Principal components allow us to reduce the number of dimensions (representative variables) of a data set and make it more manageable or interpretable while still explaining most of the original variability in the data. At the time I didn’t really understand the methodology so seek to address that retrospective learning opportunity here. I then went on to use the principal component score vectors as features for regression, this is another use for PCA. I remember using PCA for the first time as a Zoology undergraduate on the characteristics of Lavender plants either hosting a Crab Spider or not. Supporting the notion, that the knowledge of how a method works, helps to avoid misinterpretation and strengthens the conclusions one draws (see here for a more detailed discussion of PCA). The authors argue, more generally, for a careful use of the analysis tool when interpreting data. For the application of PCA to genetic data, take a look at the paper by Reich et al 2008. Principal Component Analysis (PCA) is a common technique for finding patterns in data of high dimension. Instead it may be preferred to have a “sorta” understanding of what Principal Components are and when we should use them (Dennett, 2013).
#Pca column 64bit 4shared full#
It can be hard staying on top of all this esoteric knowledge and having a full understanding. This is challenging in contrast to supervised learning, where there is a simple goal for the analysis, here there is no way to check our work because we don’t know the true answer (we have no training set to compare our predictions against). Instead of attempting to make predictions we can try to make sense of the data using unsupervised learning techniques. Sometimes one gets handed so much data you don’t know where to begin! You might not even have an associated response variable to complement the hundreds of explanatory variables provided.

